The process is quite rapid and occurs without many mistakes.ĭNA replication employs a large number of proteins and enzymes, each of which plays a critical role during the process. This means that approximately 1000 nucleotides are added per second. coli has 4.6 million base pairs in a single circular chromosome and all of it gets replicated in approximately 42 minutes, starting from a single origin of replication and proceeding around the circle in both directions. Okazaki fragments: Fragments of newly synthesized DNA formed on the lagging strand.ĭNA ligase: An enzyme that seals gaps between Okazaki fragments on the lagging strand.ĭNA proofreading: An enzymatic function that some polymerases have to recognize and repair incorrectly added nucleotides during replication.DNA replication has been extremely well studied in prokaryotes primarily because of the small size of the genome and the mutants that are available. Lagging strand: One of the two newly synthesized DNA strands that must be synthesized in the opposite direction from the replication fork. Primase: An enzyme that synthesizes RNA primers complementary to the DNA strand.ĭNA polymerase III: An enzyme that extends the RNA primers by adding nucleotides in the 5′ to 3′ direction the main factor that synthesizes new DNA.Įxonuclease: An enzyme that removes nucleotides from the end of a DNA or RNA molecule.ĭNA polymerase I: An enzyme that removes the RNA primers and replaces them with newly synthesized DNA. Sliding clamp: An enzyme that holds the DNA polymerase in place when nucleotides are being added. Supercoiling: Over or underwinding of the DNA helix that occurs as the double-stranded helix is separated during replication this puts strain on the DNA and can inhibit replication. Topoisomerase: An enzyme that functions ahead of the replication fork to prevent supercoiling of the DNA by introducing breaks and then sealing them. 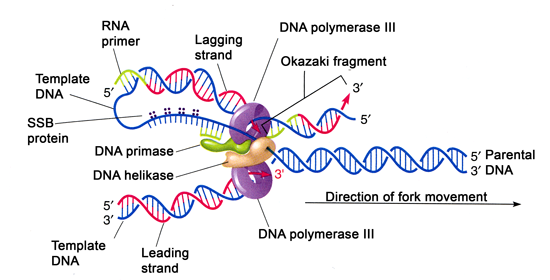
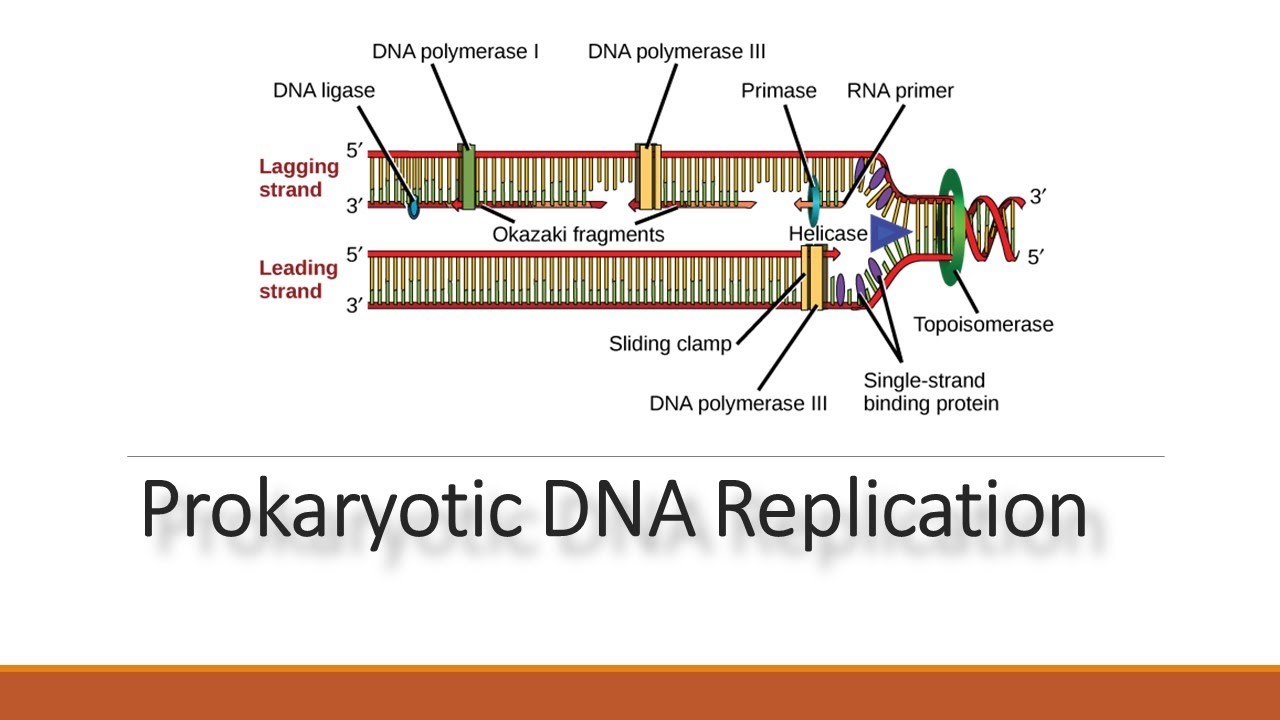
Replication fork: A Y-shaped structure that forms as the double-stranded DNA helix is separated by helicase. Single-strand binding proteins (SSBs): An enzyme that coats the separated DNA strands around the replication fork to prevent rewinding of the DNA. Helicase: An enzyme that unwinds the DNA helix ahead of the replication machinery by breaking hydrogen bonds between nucleotide base pairs. 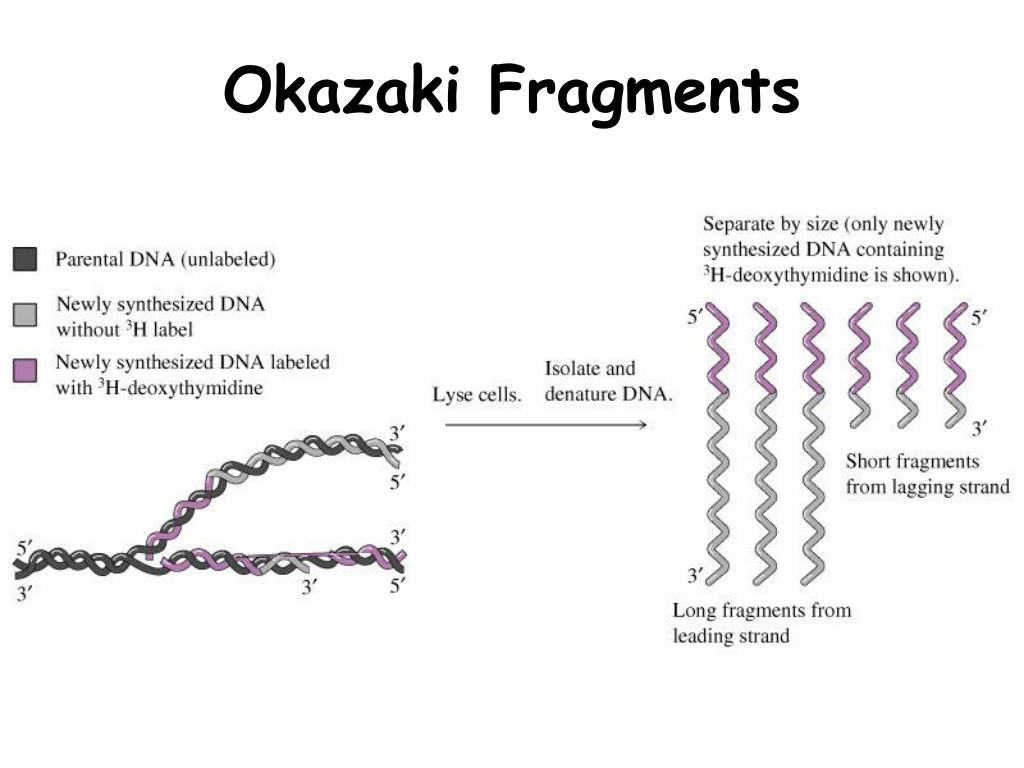
Prokaryote: Single-celled organisms without membrane-bound organelles.Įukaryote: Organisms with membrane-bound organelles. The basics of DNA replication are similar in prokaryotes and eukaryotes, but eukaryotes have many more enzymes involved.Ĭatalysis: The speeding up of a chemical reaction, in this case by an enzyme.ĭNA replication: The process by which DNA is copied.The enzymes involved in the replication of prokaryotic DNA are DNA polymerase I to III, helicase, ligase, primase, sliding clamp, topoisomerase, and single-strand binding proteins (SSBs).In addition, a separate RNase enzyme helps in the removal of RNA primers instead of DNA polymerase I. For instance, there are five human DNA polymerases with important roles in replication. However, a single enzyme in prokaryotes may be represented by multiple enzymes in eukaryotes. The enzymes involved in eukaryotic replication are similar to those involved in prokaryotic replication. There is a third polymerase involved, called DNA polymerase II, which has a DNA proofreading ability and serves a repair function. The gaps between the DNA fragments on the lagging strand, known as Okazaki fragments, are sealed by DNA ligase. RNA primers are removed (by exonuclease activity) and replaced with newly synthesized DNA by DNA polymerase I. DNA polymerase III extends the primers, adding nucleotides in the 5′ to 3′ direction, to make the bulk of the new DNA. Primase synthesizes RNA primers complementary to the DNA strand. The sliding clamp helps to hold the DNA polymerase in place when nucleotides are being added. Topoisomerase works at the region ahead of the replication fork to prevent supercoiling by introducing breaks in the DNA and then resealing them. Single-strand binding proteins (SSBs) coat the DNA around the replication fork to prevent rewinding of the DNA. Helicase opens up the DNA at the replication fork by breaking hydrogen bonds between the nucleotide base pairs.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
